To achieve the world's highest accuracy and speed, PEZY Computing is continuing to develop a whole human genome analysis system.
1. Overview and Features:
ZettaScaler-3.0 Server Unit provides ultra-fast secondary genomic analysis of whole human genome sequencing data with high accuracy. The server equips with four PEZY-SC3 processors, which were originally designed and developed by PEZY Computing group.
・A single server unit can process about 100 human whole genome data (equivalent to 100 Gbp/sample) per day.
・It provides results with extremely high analysis accuracy without sacrificing accuracy for speed.
Example: SNP F value: approx. 0.999, INDEL F value: approx. 0.996, processing time: approx. 15 min (100Gbp conversion)
(Genome In A Bottle, HG001 benchmark ver. 3.3.2 was used for the evaluation.)

2. Analysis Workflow:From FASTQ file input to VCF file generation
[FASTQ] → Alignment → Coordinate Sorting → Mark Duplicates → Base Quality Score Recalibration & Apply BQSR → HaplotypeCaller→ [VCF]
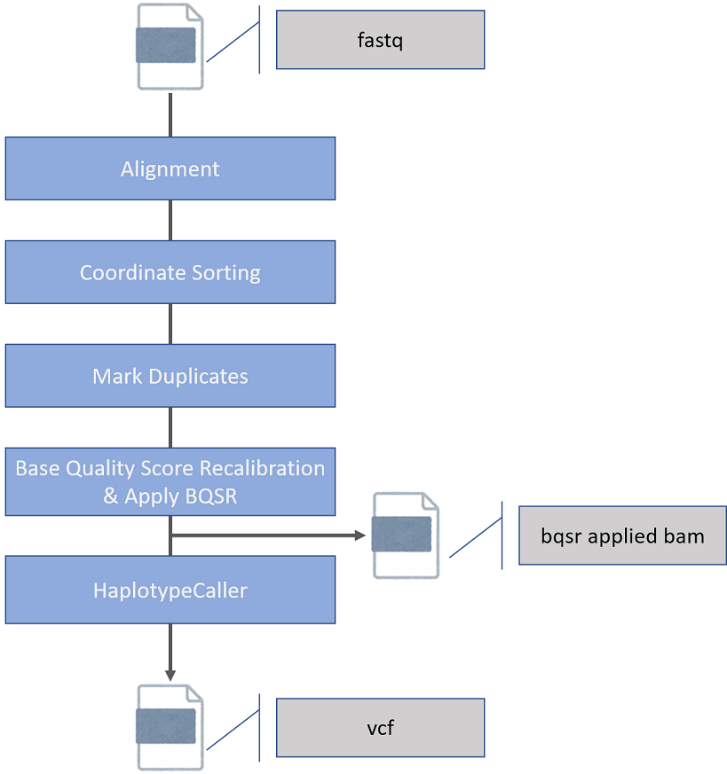
Accelerated software used in the GATK Best Practice pipeline, the most commonly used in human genome analysis, completes to variant calls in less than 15 minutes when 100 Gbase FASTQ is used as input.
The following improvements have been made to increase speed
1) Acceleration by PEZY-SC3 porting
2) Intermediate files made on-memory
3) Optimization of CPU processing
We are not only increasing speed, but also improving accuracy.
1) BWA MEM has been implemented with options to improve accuracy.
2) Highly accurate probabilistic model implemented in GATK 4.2 is implemented.
3. ZettaScaler-3.0 Server Unit specification
baseboard: ExaScaler EPX-BASE2 x 1 CPU: AMD EPYC x 1
accelerator:PEZY Computing MOD-SC3H (PCIe Gen4x16 bus) x max 4 modules
main memory:DDR4 ECC Registered 3200MHz SDRAM 1TB (max 2TB)
storage:M.2 NVMe SSD 2TB x 4
OS:AlmaLinux 9
4. PzBWA-MEM
PzBWA-MEM is a fast alignment software based on BWA-MEM version 0.7.17 (r1198) with improvements by PEZY Computing.
1) Acceleration of alignment process by PEZY-SC3
2) Optimization of the number of pipeline stages and pipeline structure for faster processing
3) Fast query data loading by optimizing Fastq loading
4) Faster post-processing by making the output on-memory
5) Add options to adjust scores, etc. (By default, it works the same as BWA-MEM.)
6) Lift-over function of alignment to Alternate contig.
7) Sensitivity improvement using information on known mutations
5. PzHaplotypeCaller
PzHaplotypeCaller is a software for genome mutation analysis based on HaplotypeCaller of GATK 4.2.0.0, which has been made faster and more accurate by PEZY Computing.
The improvements are as follows
1) Full scratch in C++ based on GATK4.2.0.0 HaplotypeCaller
2) Accelerated processing using PEZY-SC3
3) Optimization of CPU processing for higher speed
4) Probabilistic model implemented in GATK 4.2
・Foreign Read Detection (FRD)
・Base Quality Dropout (BQD)
・DragSTR
